The Second Genome Nobody Talks About (And the Tool I Built to Fix It)
This one is a little close to me — because I spent nearly a year working on it.
It was March 2024. I had just received the Khorana Scholarship and was looking for labs in the US to join for the summer. I came across the Kundu Lab at St. Jude Children’s Research Hospital in Memphis, and their work on the role of mitochondrial DNA mutations in cancer. Instantly, I was captivated. I wanted to go deeper. Thankfully, everything worked out and by May, I had landed in Memphis.
Today’s paper: “MitoEdit: A pipeline for optimizing mtDNA base editing and predicting bystander effects” — Shah, McCastlain et al. (2025)
Your Cells Have a Secret Genome
When you think of DNA, most people picture the nucleus. But the mitochondria — yes, the powerhouse of the cell — also has its own separate DNA (mtDNA).
Here’s why. Recall the endosymbiont theory: millions of years ago, a eukaryote engulfed a prokaryotic bacterium. Instead of digesting it, the two formed a symbiotic relationship — and that bacterium never left. We now call it the mitochondria. The DNA that was inside that original bacterium survived too, and we now know it as mtDNA.
Here’s the cool part: we inherit all our mitochondria only from our mothers. Every mitochondrion in your body traces back through an unbroken maternal line — all the way to a single woman scientists have nicknamed “Mitochondrial Eve.” This makes these organelles even more exciting about.
When the Powerhouse Gets a ‘Typo’ Error
Before I joined the lab, the team had already discovered a bunch of mtDNA mutations in pediatric leukemia patients — which is part of what pulled me in. And this isn’t unusual. Maternally inherited mtDNA mutations have been linked to everything from rare blindness conditions to muscle disorders, neurological diseases, and cancer. The list is long and growing.
The problem? For decades, we’ve almost entirely focused on the nuclear genome (nDNA) and largely ignored mtDNA mainly because studying and fixing mutations in the mitochondria is genuinely hard.
Why Not Just Use CRISPR?
Fair question. CRISPR is the “molecular scissors” everyone talks about, so why not just use it to fix the mutation?
Two reasons.
First, it’s incredibly difficult to get the guide RNA (the GPS that tells CRISPR where to cut) into the mitochondria. Think of it like trying to deliver a letter to a building that has no mailbox. There’s just no import system for RNA in the mitochondria. The guide RNA rarely makes it inside.
Second, CRISPR works by cutting DNA and relying on the cell’s repair machinery to introduce the change. But mitochondrial DNA repair is so poor that if you cut mtDNA, it doesn’t repair, it simply degrades. You’ve destroyed what you were trying to fix.
Enter the ‘Pencil’ (aka DdCBEs)
In 2020, the Liu Lab at Harvard introduced a specialized editing system built specifically for the mitochondria: DdCBEs (DddA-derived cytosine base editors).
Here’s how it works. A bacterial toxin called DddA acts as the actual editor — it changes one DNA letter at a time (C to T, specifically). It’s guided to the right spot on the mtDNA by protein structures called TALE domains — think of them as the address label on a package, telling the editor exactly which part of the mitochondrial genome to go to.
Unlike CRISPR, there’s no cutting. No relying on broken repair machinery. Just a pencil, quietly changing one letter at a time.
The Pencil Has Fat Fingers
Here’s the catch.
The TALE domains aren’t perfectly precise. When the editor arrives at the target and starts changing the intended letter, it sometimes accidentally changes the nearby letters too. These are called bystander edits, and the frustrating part is that you don’t know which ones will happen until after you’ve run the experiment.
It’s like autocorrect changing words you didn’t ask it to change but you only find out after you’ve already sent the message.
The old workflow: build multiple versions of the editor construct, run each one in the lab, test which version causes the least unintended changes. Time-consuming. Expensive. Slow.
The Spell-Checker Tool : MitoEdit
This is where MitoEdit comes in.
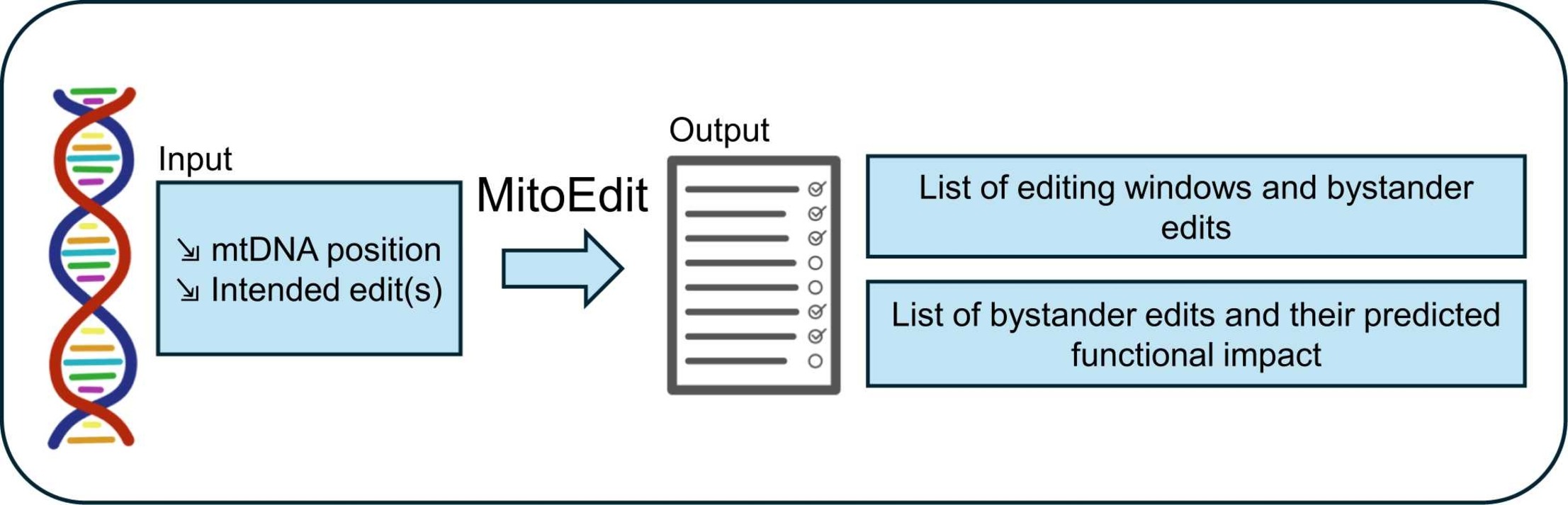
Instead of finding out after the experiment which letters got accidentally changed, MitoEdit tells you before. You input the mutation you want to introduce, and the tool predicts which nearby letters are likely to get accidentally edited.
You pick the editing window with the least mess. Then you go to the lab. Fewer constructs to build. Less time wasted.
We validated MitoEdit against publicly available data and found predictions are pretty accurate for most cases. It does start to underestimate a bit when there are many bystander edits happening in a single window — something we actively tried to improve, but we still don’t have an answer.
The tool is available at mitoedit.stjude.org.
The Bigger Picture!
Mitochondrial research has gained serious ground over the past few years. Mutations in mtDNA have been linked to everything from rare inherited conditions to cancer — including the pediatric leukemia cases that first brought me to Memphis. And yet, for most of these mutations, we still don’t have therapies.
Tools like MitoEdit are a small but real step toward changing that — making it faster to model these diseases, study them properly, and eventually treat them. The next decade of mitochondrial biology is going to be exciting.
I don’t know how big a role MitoEdit will play in that story. But building it feels like a small piece of the puzzle.
Want to discuss this paper? Have questions? Reach out!
📧 Email: devansh.shah@iitb.ac.in
Feel free to share your thoughts, corrections, or follow-up questions. We’d love to hear from you!
References
- Shah, D., McCastlain, K., Chang, T. C., Zhu, X., Wu, G., & Kundu, M. (2025). MitoEdit: A pipeline for optimizing mtDNA base editing and predicting bystander effects. Computational and Structural Biotechnology Journal, 27, 1673–1676. https://doi.org/10.1016/j.csbj.2025.04.027
- Mok, B. Y., et al. (2020). A bacterial cytidine deaminase toxin enables CRISPR-free mitochondrial base editing. Nature, 583, 631–637.
- MitoEdit Web Interface: https://mitoedit.stjude.org/
- MitoEdit GitHub: https://github.com/Kundu-Lab/mitoedit